|
x |
x |
|
 |
 |
|
INFECTIOUS
DISEASE |
BACTERIOLOGY |
IMMUNOLOGY |
MYCOLOGY |
PARASITOLOGY |
VIROLOGY |
|
|
BACTERIOLOGY - CHAPTER EIGHT
EXCHANGE OF GENETIC INFORMATION
Gene Mayer, PhD
Professor Emeritus
University of South Carolina School of Medicine
Columbia
South Carolina
|
|
SPANISH |
|
|
|
TURKISH |
Let us know what you think
FEEDBACK |
|
SEARCH |
|
|
|

 |
|
Logo image © Jeffrey
Nelson, Rush University, Chicago, Illinois and
The MicrobeLibrary |
|
|
|
TEACHING OBJECTIVES
To explain the mechanisms of gene transfer in
bacteria
To describe the nature of transposable genetic elements and plasmids
To discuss the significance of gene transfer, transposable
genetic elements and plasmids |
INTRODUCTION
In bacterial populations mutations are constantly arising due
to errors made during replication. If there is any selective advantage for a
particular mutation (e.g. antibiotic resistance), the mutant will
quickly become the major component of the population due to the rapid growth
rate of bacteria. In addition, since bacteria are haploid organisms, even
mutations that might normally be recessive will be expressed. Thus, mutations
in bacterial populations can pose a problem in the treatment of bacterial
infections. Not only are mutations a problem, bacteria have mechanisms by
which genes can be transferred to other bacteria. Thus, a mutation arising in
one cell can be passed on to other cells.
Gene transfer in bacteria is unidirectional from a donor cell
to a recipient cell and the donor usually gives only a small part of its DNA
to the recipient. Thus, complete zygotes are not formed; rather, partial
zygotes (merozygotes) are formed.
Bacterial genes are usually transferred to members of the same
species but occasionally transfer to other species can also occur. Figure 1
illustrates gene transfers that have been shown to occur between different
species of bacteria.
|
KEY WORDS
Merozygote
Transformation
Competence
Homologous
recombination
Transduction
Generalized
transduction
Specialized
transduction
Lysogenic conversion
Conjugation
F/sex
pilus
Replicon
F+
F-
Hfr
F'
Transposable genetic
element
Insertion
sequence
Transposon
Site-specific recombination
Phase
variation
Plasmid, Conjugative
plasmid
Nonconjugative plasmid
R
factor
RTF
R determinant |
GENE TRANSFER MECHANISMS IN BACTERIA
Transformation
Transformation is gene transfer
resulting from the uptake by a recipient cell of naked DNA from a donor
cell. Certain bacteria (e.g. Bacillus, Haemophilus, Neisseria,
Pneumococcus) can take up DNA from the environment and the DNA that is taken
up can be incorporated into the recipient's chromosome.
Factors affecting transformation
a. DNA size state
Double stranded DNA of
at least 5 X 105 daltons works best. Thus, transformation is
sensitive to nucleases in the environment.
b. Competence of the recipient
Some
bacteria are able to take up DNA naturally. However, these bacteria only
take up DNA a particular time in their growth cycle when they produce a
specific protein called a competence factor. At this stage the bacteria
are said to be competent. Other bacteria are not able to take up DNA
naturally. However, in these bacteria competence can be induced in
vitro by treatment with chemicals (e.g. CaCl2).
Steps in transformation
a. Uptake of DNA
Uptake of DNA by Gram+
and Gram- bacteria differs. In Gram + bacteria the DNA is taken up as a
single stranded molecule and the complementary strand is made in the
recipient. In contrast, Gram- bacteria take up double stranded DNA.
b. Legitimate/Homologous/General Recombination
After the donor DNA is taken up, a reciprocal recombination event
occurs between the chromosome and the donor DNA. This recombination
requires homology between the donor DNA and the chromosome and results
in the substitution of DNA between the recipient and the donor as
illustrated in Figure 2.
|
 E. coli (rod prokaryote) strains undergoing conjugation. One strain has fimbriae
©
Dr Dennis
Kunkel, University of Hawaii. Used with permission
E. coli (rod prokaryote) strains undergoing conjugation. One strain has fimbriae
©
Dr Dennis
Kunkel, University of Hawaii. Used with permission
 Figure 1 Gene transfers that have been shown to occur between different
species of bacteria
Figure 1 Gene transfers that have been shown to occur between different
species of bacteria
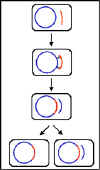 Figure 2 General recombination. Donor DNA is
shown in red and recipient DNA in blue
Figure 2 General recombination. Donor DNA is
shown in red and recipient DNA in blue
 Fig. 3 The mechanism of generalized transduction
Fig. 3 The mechanism of generalized transduction |
Recombination requires the bacterial recombination genes
(recA, B and C) and homology between the DNA's involved. This type of
recombination is called legitimate or homologous or general
recombination. Because of the requirement for homology between the
donor and host DNA, only DNA from closely related bacteria would be
expected to successfully transform, although in rare instances gene
transfer between distantly related bacteria has been shown to occur.
Significance
Transformation occurs in nature
and it can lead to increased virulence. In addition transformation is
widely used in recombinant DNA technology.
Transduction
Transduction is the transfer of
genetic information from a donor to a recipient by way of a bacteriophage.
The phage coat protects the DNA in the environment so that transduction,
unlike transformation, is not affected by nucleases in the environment. Not
all phages can mediate transduction. In most cases gene transfer is between
members of the same bacterial species. However, if a particular phage has a
wide host range then transfer between species can occur. The ability of a
phage to mediated transduction is related to the life cycle of the phage.
Types of Transduction
a. Generalized Transduction - Generalized
transduction is transduction in which potentially any bacterial gene
from the donor can be transferred to the recipient. The mechanism of
generalized transduction is illustrated in Figure 3.
Phages that mediate generalized transduction generally
breakdown host DNA into smaller pieces and package their DNA into the
phage particle by a "head-full" mechanism. Occasionally one of
the pieces of host DNA is randomly packaged into a phage coat. Thus, any
donor gene can be potentially transferred but only enough DNA as can fit
into a phage head can be transferred. If a recipient cell is infected by
a phage that contains donor DNA, donor DNA enters the recipient. In the
recipient a generalized recombination event can occur which substitutes
the donor DNA and recipient DNA (See Figure 2).
|
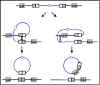 Figure 4 The mechanism of specialized transduction
Figure 4 The mechanism of specialized transduction |
b. Specialized transduction - Specialized
transduction is transduction in which only certain donor genes can be
transferred to the recipient. Different phages may transfer different
genes but an individual phage can only transfer certain genes.
Specialized transduction is mediated by lysogenic or temperate phage and
the genes that get transferred will depend on where the prophage has
inserted in the chromosome. The mechanism of specialized transduction is
illustrated in Figure 4.
During excision of the
prophage, occasionally an error occurs
where some of the host DNA is excised with the phage DNA. Only host DNA on
either side of where the prophage has inserted can be transferred (i.e.
specialized transduction). After replication and release of phage and
infection of a recipient, lysogenization of recipient can occur resulting in
the stable transfer of donor genes. The recipient will now have two copies of
the gene(s) that were transferred. Legitimate recombination between the donor
and recipient genes is also possible.
Significance
Lysogenic (phage) conversion
occurs in nature and is the source of virulent strains of bacteria.
|
|
MOVIE
Conjugation
High
resolution
Low resolution
© Mondo Media, San Francisco, Calif., USA and
and The
MicrobeLibrary
This video clip demonstrates the process of conjugation. First, two bacteria combine via a sex pilus. Next, one strand of the plasmid is transferred to the attached cell. Note that the original plasmid is not lost from the first cell. Finally, each cell immediately duplicates the single strand so that both bacteria have
a copy of the double stranded plasmid
|
Conjugation
Transfer of DNA from a donor to a
recipient by direct physical contact between the cells. In bacteria there
are two mating types a donor (male) and a recipient (female) and the
direction of transfer of genetic material is one way; DNA is transferred
from a donor to a recipient.
Mating types in bacteria
a. Donor
The ability of a bacterium to be a
donor is a consequence of the presence in the cell of an extra piece of DNA
called the F factor or fertility factor or sex factor.
The F factor is a circular piece of DNA that can replicate autonomously in
the cell; it is an independent replicon. Extrachromosomal pieces of
DNA that can replicate autonomously are given the general name of plasmids.
The F factor has genes on it that are needed for its replication and for its
ability to transfer DNA to a recipient. One of the things the F factor codes
for is the ability to produce a sex pilus (F pilus) on the surface of
the bacterium. This pilus is important in the conjugation process. The F
factor is not the only plasmid that can mediated conjugation but it is
generally used as the model.
b. Recipient
The ability to act as a
recipient is a consequence of the lack of the F factor.
|
| |
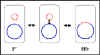 a
a
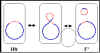 b Figure 5 Physiological states of F factor
b Figure 5 Physiological states of F factor
|
Physiological states of the F factor
a. Autonomous (F+)
In this state
the F factor carries only those genes necessary for its replication and for
DNA transfer. There are no chromosomal genes associated with the F factor in
F+ strains.
In crosses of the type F+ X F- the F-
becomes F+ while F+ remains F+. Thus, the F
factor is infectious. In addition, there is only low level transfer of
chromosomal genes.
b. Integrated
(Hfr)
In this state the
F factor has integrated into the bacterial chromosome via a
recombination event as illustrated in the Figure 5a
In crosses of the type Hfr X F- the F-
rarely becomes Hfr and Hfr remains Hfr. In addition, there is a high
frequency of transfer of donor chromosomal genes.
|
| |
c. Autonomous with chromosomal genes (F')
In
this state the F factor is autonomous but it now carries some chromosomal
genes. F' factors are produced by excision of the F factor from an Hfr, as
illustrated in Figure 5b. Occasionally, when the F factor is excising from
the Hfr chromosome, donor genes on either side of the F factor can be
excised with the F factor generating an F'. F' factors are named depending
on the chromosomal genes that they carry.
In crosses of the type F' X F- the F-
becomes F' while F' remains F'. In addition there is high frequency of
transfer of those chromosomal genes on the F' and low frequency transfer of
other donor chromosomal genes.
|
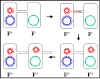 Figure 6 Mechanism of F+ x F- crosses
Figure 6 Mechanism of F+ x F- crosses |
Mechanism of conjugation
a. F+ X F- crosses
(Figure 6)
i) Pair formation
The tip of the sex pilus comes in
contact with the recipient and a conjugation bridge is formed
between the two cells. It is through this bridge that the DNA will pass
from the donor to the recipient. Thus, the DNA is protected from
environmental nucleases. The mating pairs can be separated by shear forces
and conjugation can be interrupted. Consequently, the mating pairs remain
associated for only a short time.
ii) DNA transfer
The plasmid DNA is nicked at a specific
site called the origin of transfer and is replicated by a rolling
circle mechanism. A single strand of DNA passes through the conjugation
bridge and enters the recipient where the second strand is replicated.
iii) This process explains the characteristics of F+
X F- crosses. The recipient becomes F+, the donor
remains F+ and there is low frequency of transfer of donor
chromosomal genes. Indeed, as depicted in Figure 7 there is no transfer of
donor chromosomal genes. In practice however, there is a low level of
transfer of donor chromosomal genes in such crosses.
|
|
ANIMATION
Mating of F+ and F- Bacterial Strains
© Thomas M. Terry, University of Connecticut, Storrs, Conn., USA
and
The
MicrobeLibrary
The F plasmid is a self-transmissible plasmid found in some strains of E. coli. Cells that possess one or more copies of the F plasmid are called F+; cells lacking the F plasmid are called F-. The animation illustrates several stages in the transfer of the F plasmid from F+ to F- cells.
|
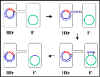 Figure 7 Mechanism of Hfr x F-
crosses
Figure 7 Mechanism of Hfr x F-
crosses |
b. Hfr X F- crosses (Figure 7)
i) Pair Formation
ii) DNA transfer
The DNA is nicked at the origin of
transfer and is replicated by a rolling circle mechanism. But the DNA that
is transferred first is the chromosome. Depending upon where in the
chromosome the F factor has integrated and in what orientation, different
chromosomal genes will be transferred at different times. However, the
relative order and distances of the genes will always remain the same.
Only when the entire chromosome is transferred will the F
factor be transferred. Since shearing forces separate the mating pairs it is
rare that the entire chromosome will be transferred. Thus, the recipient
does not receive the F factor in a Hfr X F- cross.
iii) Legitimate recombination
Recombination between the
transferred DNA and the chromosome results in the exchange of genetic
material between the donor and recipient.
iv) This mechanism explains the characteristics of Hfr X F-
crosses. The recipient remains F-, the donor remains Hfr and there is a high
frequency of transfer of donor chromosomal genes.
|
|
ANIMATION
Mating of Hfr and F- Bacterial Strains
© Thomas M. Terry, University of Connecticut, Storrs, Conn., USA
and
The
MicrobeLibrary
|
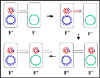 Figure 8 The mechanism of F" x F- crosses
Figure 8 The mechanism of F" x F- crosses |
c. F' X F- crosses
(Figure 8)
i) Pair formation
ii) DNA transfer
This process is similar to F+ X F- crosses. However,
since the F' has some chromosomal genes on it these will also be
transferred.
iii) Homologous
recombination is not necessary although it may occur.
iv) This mechanism
explains the characteristics of F' X F-
crosses. The F- becomes F', the F' remains F' and the is high frequency
transfer of donor genes on the F' but low frequency transfer of other donor
chromosomal genes.
Significance
Among the Gram negative bacteria this is the major way that bacterial genes
are transferred. Transfer can occur between different species of bacteria.
Transfer of multiple antibiotic resistance by conjugation has become a major
problem in the treatment of certain bacterial diseases. Since the recipient
cell becomes a donor after transfer of a plasmid it is easy to see why an
antibiotic resistance gene carried on a plasmid can quickly convert a
sensitive population of cells to a resistant one.
Gram positive
bacteria also have plasmids that carry multiple antibiotic resistance genes,
in some cases these plasmids are transferred by conjugation while in others
they are transferred by transduction. The mechanism of conjugation in Gram +
bacteria is different than that for Gram -. In Gram + bacteria the donor
makes an adhesive material which causes aggregation with the recipient and
the DNA is transferred.
|
| |
TRANSPOSABLE GENETIC ELEMENTS
Transposable Genetic Elements
Transposable
genetic elements are segments of DNA that have the capacity to move from one
location to another (i.e. jumping genes).
Properties of Transposable Genetic Elements
Random movement
Transposable genetic elements
can move from any DNA molecule to any DNA other molecule or even to
another location on the same molecule. The movement is not totally random;
there are preferred sites in a DNA molecule at which the transposable
genetic element will insert.
Not capable of self
replication
The
transposable genetic elements do not exist autonomously (exception - some
transposable phages) and thus, to be replicated they must be a part of
some other replicon.
Transposition mediated by
site-specific recombination
Transposition requires little or no homology between
the current location and the new site. The transposition event is mediated
by a transposase coded for by the transposable genetic
element. Recombination that does not require homology between the
recombining molecules is called site-specific or illegitimate or
nonhomologous recombination.
Transposition can be
accompanied by duplication
In many instances transposition of the transposable genetic element
results in removal of the element from the original site and insertion at
a new site. However, in some cases the transposition event is accompanied
by the duplication of the transposable genetic element. One copy remains
at the original site and the other is transposed to the new site.
|
 Figure 9 Structure of transposable genetic elements
Figure 9 Structure of transposable genetic elements |
Types of Transposable Genetic Elements
Insertion sequences (IS)
Insertion sequences
are transposable genetic elements that carry no known genes except those
that are required for transposition.
a. Nomenclature
Insertion sequences are
given the designation IS followed by a number. e.g. IS1
b. Structure (Figure 9)
Insertion sequences are small stretches of DNA that have
at their ends repeated sequences, which are involved in transposition.
In between the terminal repeated sequences there are genes involved in
transposition and sequences that can control the expression of the genes
but no other nonessential genes are present.
c. Importance
i) Mutation
The introduction of an insertion sequence
into a bacterial gene will result in the inactivation of the gene.
ii) Plasmid insertion into
chromosomes
The sites at
which plasmids insert into the bacterial chromosome are at or near
insertion sequence in the chromosome.
iii) Phase Variation
The flagellar antigens are one of
the main antigens to which the immune response is directed in our
attempt to fight off a bacterial infection. In Salmonella there are two
genes which code for two antigenically different flagellar antigens. The
expression of these genes is regulated by an insertion sequences. In one
orientation one of the genes is active while in the other orientation
the other flagellar gene is active. Thus, Salmonella can change their
flagella in response to the immune systems' attack. Phase variation is
not unique to Salmonella flagellar antigens. It is also seen with other
bacterial surface antigens. Also the mechanism of phase variation may
differ in different species of bacteria (e.g. Neisseria;
transformation).
Transposons
(Tn)
Transposons are
transposable genetic elements that carry one or more other genes in
addition to those which are essential for transposition.
|
 Figure 10 Transposon structure
Figure 10 Transposon structure |
a. Nomenclature
Transposons are given
the designation Tn followed by a number.
b. Structure
The structure of a transposon is similar to that of an insertion sequence. The extra genes
are located between the terminal repeated sequences. In some instances
(composite transposons) the terminal repeated sequences are actually
insertion sequences. (See Figure 10).
c. Importance
Many antibiotic resistance
genes are located on transposons. Since transposons can jump from one
DNA molecule to another, these antibiotic resistance transposons are a
major factor in the development of plasmids which can confer multiple
drug resistance on a bacterium harboring such a plasmid. These multiple
drug resistance plasmids have become a major medical problem because the
indiscriminate use of antibiotics have provided a selective advantage
for bacteria harboring these plasmids.
|
| |
PLASMIDS
Definition
Plasmids are extrachromosomal genetic elements capable of autonomous
replication. An episome is a plasmid that can integrate into the
bacterial chromosome.
Classification
of Plasmids
Transfer
properties
a. Conjugative
plasmids
Conjugative plasmids are those that mediated
conjugation. These plasmids are usually large and have all the genes
necessary for autonomous replication and for transfer of DNA to a
recipient (e.g. genes for sex pilus).
b. Non-conjugative
plasmids
Non-conjugative plasmids are those that cannot mediate
conjugation. They are usually smaller than conjugative plasmids and they
lack one or more of the genes needed for transfer of DNA. A
non-conjugative plasmid can be transferred by conjugation if the cell
also harbors a conjugative plasmid.
Phenotypic
effects
a. Fertility plasmid (F factor)
b. Bacteriocinogenic
plasmids
These plasmids have genes which code for substances that
kill other bacteria. These substances are called bacteriocins or
colicins.
c. Resistance
plasmids 7 factors)
These plasmids carry antibiotic resistance
genes.
i) Origin -
The origin of the R factors is not known. It is likely that they
evolved for other purposes and the advent of the antibiotic age
provided a selective advantage for their wide-spread dissemination.
|
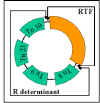 Figure 11 R plasmid structure
Figure 11 R plasmid structure |
ii) Structure - R plasmids are conjugative plasmids in which the
genes for replication and transfer are located on one part of the R factor and
the resistance genes are located on another part as illustrated in Figure 11.
RTF (Resistance
Transfer Factor)
Carries the transfer
genes.
R determinant
Carries the resistance genes. The resistance
genes are often parts of transposons.
Mode of action of resistance genes
a) Modification (detoxification) of antibiotic -
e.g.
β-lactamase
b) Alteration of target site -
e.g. Streptomycin
resistance
c) Alteration of uptake - Tetracycline resistance
d) Replacement of sensitive pathway -
e.g. new folic
acid pathway for resistance to sulfa drugs
|
|
|
 Return to the Bacteriology Section of the
Microbiology and Immunology On-line Textbook Return to the Bacteriology Section of the
Microbiology and Immunology On-line Textbook
This page last changed on
Saturday, February 27, 2016
Page maintained by
Richard Hunt
|

 E. coli (rod prokaryote) strains undergoing conjugation. One strain has fimbriae
©
Dr Dennis
Kunkel, University of Hawaii. Used with permission
E. coli (rod prokaryote) strains undergoing conjugation. One strain has fimbriae
©
Dr Dennis
Kunkel, University of Hawaii. Used with permission
 Figure 4 The mechanism of specialized transduction
Figure 4 The mechanism of specialized transduction a
a
 Figure 6 Mechanism of F+ x F- crosses
Figure 6 Mechanism of F+ x F- crosses Figure 7 Mechanism of Hfr x F-
crosses
Figure 7 Mechanism of Hfr x F-
crosses Figure 8 The mechanism of F" x F- crosses
Figure 8 The mechanism of F" x F- crosses Figure 11 R plasmid structure
Figure 11 R plasmid structure

