|
x |
x |
|
 |
 |
|
INFECTIOUS
DISEASE |
BACTERIOLOGY |
IMMUNOLOGY |
MYCOLOGY |
PARASITOLOGY |
VIROLOGY |
|
TURKISH
|
VIROLOGY - CHAPTER
FOUR
RNA VIRUS REPLICATION
STRATEGIES
Dr Margaret Hunt
Professor Emerita
Department of Pathology, Microbiology and Immunology
University of South Carolina School of Medicine
|
|
En
Español |
|
EN FRANCAIS |
|
NË SHQIPTARE |
Let us know what you think
FEEDBACK |
|
SEARCH |
|
|

 |
|
|
|
|
|
TEACHING OBJECTIVES
Descriptive analysis of the replicative
strategies employed by animal RNA viruses
Identification of virus prototypes
associated with different RNA virus replication schemes
 Structure of Polio Type 1 Mahoney. X-ray data from Hogle et al.(Harvard Univ.), PDB entry 2PLV, rendered with GRASP (A.Nicholls, Columbia Univ.). Courtesy of
Dr Sgro
and the
Institute for Molecular
Virology, Univ. of Wisconsin (used with permission)
Structure of Polio Type 1 Mahoney. X-ray data from Hogle et al.(Harvard Univ.), PDB entry 2PLV, rendered with GRASP (A.Nicholls, Columbia Univ.). Courtesy of
Dr Sgro
and the
Institute for Molecular
Virology, Univ. of Wisconsin (used with permission) |
RNA VIRUS REPLICATION - GENERAL
STRATEGIES
RNA viruses that do not
have a DNA phase
Viruses that replicate via RNA intermediates need an RNA-dependent
RNA-polymerase to replicate their RNA, but animal cells do not seem to possess a
suitable enzyme. Therefore, this type of animal RNA virus needs to code for an
RNA-dependent RNA polymerase.
No viral proteins can be made until viral messenger RNA is available; thus, the
nature of the RNA in the virion affects the strategy of the virus:
Plus-stranded RNA viruses
In these viruses, the virion (genomic) RNA
is the same sense as mRNA and so functions as mRNA. This mRNA can be translated immediately upon infection
of the host cell
Examples:
Negative-stranded RNA
viruses
The virion RNA is negative sense
(complementary to mRNA) and must therefore be copied into the complementary plus-sense
mRNA before proteins can be made. Thus, besides needing to code for an
RNA-dependent RNA-polymerase, these viruses also need to package it in the
virion so that they can make mRNAs upon infecting the cell.
Examples:
Double-stranded RNA
viruses
The virion (genomic) RNA is double
stranded and so cannot function as mRNA; thus these viruses also need to package an RNA
polymerase to make their mRNA after infection of the host cell.
Example:
RNA viruses that copy
their RNA to DNA
These are the
retroviruses. In this case, their virion RNA, although plus-sense, does not function as mRNA immediately on
infection since it is not released from the capsid into the cytoplasm. Instead, it serves as a
template for reverse transcriptase and is copied into DNA. Reverse
transcriptase is not available in the cell, and so these viruses need to code for this enzyme
and package it in virions.
|
RNA VIRUSES THAT DO NOT HAVE A
DNA PHASE |
|
Genome |
RNA-dependent RNA polymerase (=transcriptase) in virion |
Infectivity of RNA |
Initial event in cell |
| Plus-stranded RNA |
No |
Infectious |
Translation |
| Negative-stranded RNA |
Yes |
Non-infectious |
Transcription |
| Double -stranded RNA |
Yes |
Non-infectious |
Transcription |
|
RETROVIRUSES |
|
Genome |
RNA-dependent RNA polymerase (=transcriptase) in
virion |
Infectivity of RNA |
Initial event in cell |
| Plus-stranded RNA |
Yes |
Non-infectious |
Reverse transcription |
THE TRANSLATION PROBLEM
Eucaryotic host cell translation
protein synthesis machinery in
general uses
monocistronic mRNAs and so there is a problem in making more than one
type of protein from a single mRNA.
RNA viruses have several solutions to this problem:
-
The virus makes multiple monocistronic
mRNAs
-
The virus makes primary transcripts
which are processed by the host splicing machinery to give more than one monocistronic RNA
-
The viral mRNA acts as a monocistronic transcript. A large polypeptide (called a polyprotein) is made which is
then cleaved into separate proteins - Thus, one initial translation product is
processed to give rise to multiple proteins. This happens, for example, in
picornaviruses
-
The viral mRNA has special features which enable ribosomes
to bind internally instead of (or as well as) at the 5’ end
GENOME SIZE OF RNA VIRUSES
RNA viruses tend to have a relatively
small genome (although
virion size may not necessarily be small). This is
probably because the lack of RNA error correction mechanisms puts a limit on the
size of RNA genomes.
The result of having a small genome is
that RNA viruses tend to code for only a few proteins. These will include a
polymerase which can copy RNA into a complementary nucleic acid (either RNA or,
as in the case of retroviruses, DNA) and a viral attachment protein.
|
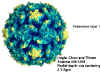 Figure 1 Polio virus © J-Y Sgro, Used with permission.
From
Virus World
Figure 1 Polio virus © J-Y Sgro, Used with permission.
From
Virus World
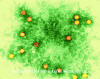 Figure 2 Polio virus x350,000 ©
Dennis Kunkel Microscopy, Inc.
Used with permission
Figure 2 Polio virus x350,000 ©
Dennis Kunkel Microscopy, Inc.
Used with permission |
POSITIVE STRAND RNA VIRUSES
Examples:
PICORNAVIRUSES
(PICORNAVIRIDAE)
Properties
These are small (28nm), naked icosahedral viruses
(figure 1) (pico=very small). The
RNA is single-stranded, plus sense, polyadenylated. It functions as mRNA immediately
upon infection
Prototype member: poliovirus (figure 1 and 2)
Adsorption and penetration
A viral protein recognizes a receptor on
the host cell
membrane (this is important in the tropism of virus).
It seems that binding to the receptor alters capsid structure in some way, a
channel forms across the cell membrane and the
RNA is released into cytoplasm. The
mRNA is now available for translation.
Synthesis of viral
proteins
Poliovirus virion RNA functions as an mRNA but does not have the methylated cap structure typical of eucaryotic mRNAs
- it has a "ribosome landing pad" (known as the internal ribosome
entry site or IRES) which
enables ribosomes to bind without having to recognize a 5' methylated cap
structure (figure 3).
Picornaviruses often interfere with host cell methylated
cap recognition. Most host cell translation is cap-dependent, so this inhibits a
lot of host protein synthesis but not viral protein synthesis - one way in which
these viruses can modify the host cell to their advantage.
|
 Figure 3
Figure 3
Structure of genomic RNA of
Picornaviridae |
The mRNA is translated into a single polypeptide (polyprotein), which is cleaved.
The cleavages occur before translation
is complete ( i.e. on the nascent (=growing) chain) and are carried out by
virally coded proteases (figure 4). Some of these proteases can work even while
part of the polyprotein.
|
 Figure 4 Adapted from Schaechter et al., Mechanisms of
Microbial Disease, 2nd Ed.
Figure 4 Adapted from Schaechter et al., Mechanisms of
Microbial Disease, 2nd Ed. |
Products of cleavage include:
An RNA polymerase (replicase)
Structural components of the virion
Proteases
|
 Figure 5
Figure 5
Replication of Picornaviridae viral
genome |
RNA replication
We now have newly made viral proteins
to support replication.
1. Viral RNA polymerase copies
plus-sense genomic RNA into complementary minus-sense RNA:
This process needs
VPg (or precursor containing VPg)
Viral
RNA polymerase (replicase)
Certain
Host proteins
VPg may act as a primer for RNA
synthesis, this would explain why it is at the 5' end of all newly synthesized
RNA molecules
2. New minus sense strands serve as
template for new plus sense strands (figure 5). Again, poliovirus RNA polymerase and VPg are
needed. VPg
is linked to the 5' ends of the new plus sense strands (again, it probably functions as a
primer).
The new plus strand has three
alternative fates:
i. It may serve as a template for more minus strands
ii. It may be packaged into progeny virions
iii. It may be translated into polyprotein (In this case VPg is usually removed prior to translation)
Assembly
When sufficient plus-sense progeny
RNA and virion proteins have accumulated, assembly begins.
Particles assemble with VPg-RNA
inside and 3 proteins in the capsid [VP0,1 and 3].
VP0 is then cleaved to VP2 and VP4 as the virions mature
and these mature virions are infectious.
Virions are released following cell
lysis. Excess capsids are formed and inclusion bodies may be seen in the
cytoplasm.
Note: The entire life cycle occurs in the cytoplasm. There is no
division into early and late gene expression
|
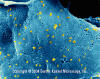 Figure 6 Rhabdovirus on a Fish Epithelial Cell
©
Dennis Kunkel Microscopy, Inc.
Used with permission
Figure 6 Rhabdovirus on a Fish Epithelial Cell
©
Dennis Kunkel Microscopy, Inc.
Used with permission |
NON-SEGMENTED NEGATIVE
STRAND VIRUSES
Examples of non-segmented negative
strand RNA viruses are:
|
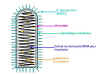 Figure 7 Structure of a typical rhabdovirus
Figure 7 Structure of a typical rhabdovirus
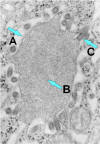 Figure 7b
Figure 7b
Rabies virus
budding from an inclusion (Negri body) into the endoplasmic reticulum in a
nerve cell. A. Negri body. B. Notice the abundant RNP in the
inclusion. C. Budding rabies virus.
CDC |
RHABDOVIRUSES
(RHABDOVIRIDAE)
Example:
Rabies virus. The most
intensively studied member is vesicular stomatitis virus.
The RNA genome:
Attachment, penetration
and uncoating
The virus adsorbs to cell surface.
G (Glycoprotein) is the attachment protein (figure 7) which binds to a receptor on
the host cell surface.
The attached virus is taken up by endocytosis.
The membrane of the virus fuses with the endosome
membrane (the acid pH of endosome is important because the G protein needs to be
exposed to acid pH before it can facilitate fusion ).
As a result of fusion of the viral membrane with the endosome membrane, the nucleocapsid is released into
cytoplasm.
Transcription
'Transcription' is used in this
context to refer to synthesis of mRNAs.
Complete uncoating of the nucleocapsid
is not necessary for transcription - the virion RNA polymerase can copy virion RNA when it is in
the nucleocapsid form. This is an advantage in that genomic RNA is therefore
somewhat protected from ribonucleases.
There is one monocistronic mRNA for
each of the five virally coded proteins (figure 8).
The mRNAs are capped, methylated,
and polyadenylated.
Since this is a cytoplasmic,
negative-sense RNA virus, the enzymes for mRNA synthesis and modification are
packaged in the virion.
Translation
Messenger RNAs are translated on host
ribosomes and all five viral proteins made at the same time.
There is no distinction between early and late functions.
|
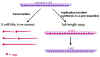 Figure 8
Figure 8
Transcription and replication of Rhabdovirus RNA |
RNA replication
RNA replication is the process by
which new copies of genome-length RNAs are made (figure 8).
RNA replication occurs in the
cytoplasm and is carried out by the viral RNA polymerase.
The full length plus strand is
coated with nucleocapsid protein as it is made (mRNAs are not coated with this
protein, which would interfere with the host protein translation machinery).
The new positive strand is copied into full
length minus strand, which is also coated with nucleocapsid protein as it is
made.
(Note: since the viral RNA
polymerase synthesizes mRNAs (transcription) and full-length RNA
(replication), it is also sometimes called a transcriptase or a replicase, such
names just focus on the different aspects of the polymerase activity.)
New negative strands may:
i. be used as templates for the
synthesis of more full length plus strands
ii. be used as templates for the synthesis of more mRNAs
iii. be packaged into virions
|
 Figure 9
Figure 9
Transport of glycoproteins from the endoplasmic reticulum to the plasma
membrane |
Assembly
The virus consists of two "modules" - the
envelope and the nucleocapsid:
Envelope
Transmembrane proteins are made on
ribosomes bound to the endoplasmic reticulum. They are
inserted into the endoplasmic reticulum membrane as they are made, glycosylated in the endoplasmic reticulum and pass
through the Golgi body where substantial modification of the carbohydrate chains
occurs. They are then transported, in vesicles, to the appropriate
cell membrane; in the case of vesicular stomatitis virus, this is the plasma
membrane (figure 9).
|
 Figure 10
Figure 10
Rhabdovirus assembly |
Nucleocapsid
Synthesis of the nucleocapsid was described above. The viral RNA
polymerase complex associates with the nucleocapsids as they are formed.
Nucleocapsids bud out through
modified areas of membrane which contain G and M proteins (figure 10). The M (matrix)
protein is involved in assembly - it interacts with patches of G in the membrane
and with nucleocapsids.
Note:
The entire life cycle occurs in the cytoplasm
RNA polymerase and RNA modification enzymes are virally-coded and
present in the virus particle (virion)
There is no division between early and late stages
|
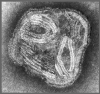 Figure 11 Paramyxovirus ©
Dr
Linda
Stannard,
University of Cape Town, South Africa
(used with permission)
Figure 11 Paramyxovirus ©
Dr
Linda
Stannard,
University of Cape Town, South Africa
(used with permission)
|
PARAMYXOVIRUSES
(PARAMYXOVIRIDAE)
Paramyxoviruses (figure 11) are
pleomorphic, that is: there are many morphological forms of the virus in a
population. They have negative-sense, non-segmented RNA and a helical nucleocapsid
(figure 12). They are enveloped, that is they are surrounded by a membrane
derived from a host cell.
The envelope contains two virally coded glycoproteins: The F protein and the
attachment protein
-
The F protein has fusion
activity
-
The attachment protein binds to
receptors on the host cell
This protein may have:
Hemagglutinating activity and
neuraminidase activity (HN protein)
or hemagglutinating activity alone (H protein)
or neither (G protein).
|
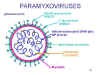 Figure 12 Structure of a typical paramyxovirus
Figure 12 Structure of a typical paramyxovirus |
|
PARAMYXOVIRUS FAMILY
SURFACE GLYCOPROTEINS |
|
GENUS
|
GLYCOPROTEIN
|
TYPICAL MEMBERS
|
|
PARAMYXOVIRUS FAMILY |
|
Paramyxovirus
|
HN, F
|
HPIV 1
HPIV 3
|
|
Rubulavirus |
HN, F |
HPIV 2
HPIV 4
mumps virus |
|
Morbillivirus
|
H, F
|
measles virus
|
|
PNEUMOVIRUS FAMILY |
|
Pneumovirus
|
G, F
|
respiratory syncytial
virus
|
|
Metapneumovirus |
G, F |
metapneumoviruses |
Hemagglutination
Hemagglutination
is easy to test for in the clinical laboratory and is used in diagnosis
Hemagglutination
involves the agglutination of red blood cells
and relies on the ability of a
virus to bind to receptors on red blood cells. Since viruses have multiple
attachment proteins per virion, they can bind to more than one red blood cell
and so they can serve to link red blood cells into a network.
Inactivated virus can still hemagglutinate as long as its attachment proteins
are intact.
If someone has antibodies to a
viral hemagglutinin, the antibodies will binds to the attachment protein and
prevent its binding to the red blood cells. The serum of that person will inhibit
the agglutination reaction by the virus to which they have antibodies - but not
by other hemagglutinating viruses. This can be used to determine which hemagglutinating
virus a person has been exposed to.
Hemadsorption
During infection, the viral attachment protein will be inserted
into the plasma membrane of the infected cell. If the viral attachment protein
can bind to red blood cells, the infected cell will bind red blood cells because
it has the viral attachment protein on its surface - this is called
hemadsorption. In the clinical laboratory, this may enable virally-infected cells to be detected at an early
stage in infection, and may allow detection of viruses which do not visibly
damage the cell.
|
 Figure 13
Figure 13
Attachment and endocytosis of paramyxoviruses |
Adsorption and penetration
The H(N)/G protein recognizes
receptors on cell surface.
The F protein facilitates fusion
between membranes at physiological pH, so although paramyxoviruses can be taken
up by endocytosis, they also often enter the cell by direct fusion with the
plasma membrane (figure 13).
Because the F protein functions at physiological pH, this can
result in
syncytia
being formed in paramyxovirus infections (see discussion of consequences of
fusion at physiological pH under DNA virus replication strategies – herpesviruses).
|
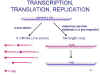 Figure 14
Figure 14
Transcription and replication of paramyxovirus RNA |
Transcription, translation
and replication of RNA
Events inside the cell are very
similar to rhabdoviruses (figure 14):
-
Viral multiplication occurs in the cytoplasm.
-
The viral RNA polymerase uses the nucleocapsid as a
template.
-
The RNA polymerase does not need a fully uncoated
nucleocapsid.
-
Viral mRNAs are transcribed; these are capped, methylated and polyadenylated.
-
Since this is a negative-strand RNA virus, RNA polymerase
and RNA modification enzymes are packaged in the virion.
-
The viral mRNAs are translated to give viral proteins.
-
There is no distinction between early and late functions in gene expression.
Viral RNA replication involves full
length plus strand synthesis. This is used as a template for full length minus
strand. Both full length strands are coated with nucleocapsid protein as they
are made (figure 14).
New full length minus strands may
serve as templates for replication, or templates for transcription, or they may
be packaged into new virions.
|
 Figure 15
Figure 15
Activation of the fusion protein by proteolytic cleavage
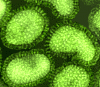 Figure 16 Orthomyxovirus (Influenza A) © Dr Linda Stannard,
University of Cape Town, South Africa
Figure 16 Orthomyxovirus (Influenza A) © Dr Linda Stannard,
University of Cape Town, South Africa
|
Assembly
Both viral glycoproteins (i.e.
attachment protein and F (fusion) protein) are translated as transmembrane
proteins and transported to the cell plasma membrane.
M (matrix) protein enables nucleocapsids to interact with the regions of the
plasma membrane which have the glycoproteins inserted.
The virus buds out through membrane.
Role of neuraminidase
In those paramyxoviruses which have
it, the neuraminidase may facilitate release. In these viruses, sialic acid
appears to be an important part of the receptor. The neuraminidase removes
sialic acid (neuraminic acid) from the cell surface. Thus, since sialic acid
will have been largely removed from the cell surface and the progeny virions,
neither will have functional receptors, so progeny virions will not stick to
each other or to the cell they have just budded out from (or any other infected
cell). They will therefore be able to diffuse away until they meet an uninfected
cell.
The neuraminidase may also help
during infection since, if the virus binds to sialic acid residues in mucus, it
would not be able to bind to a receptor on a cell and infect that cell. But if the sialic acid
in the mucus is eventually
destroyed, the virus will be freed and may then reach a receptor on the cell
surface.
Activation of the F
protein
The F protein needs to be cleaved
before it can function in facilitating fusion when the virus binds to another
cell (figure 15). This is a late event in maturation.
|
Some differences between
rhabdoviruses and paramyxoviruses |
| |
Rhabdovirus |
Paramyxovirus |
| Shape |
bullet
bacilliform |
round
pleomorphic |
| Glycoproteins |
One (has both attachment and
fusion activities) |
Two (one attachment and one
fusion) |
| Fusion pH |
acidic |
neutral
physiological |
|
|
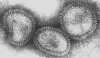 Figure 17 Orthomyxovirus (Influenza A) © Dr Linda Stannard,
University of Cape Town, South Africa
Figure 17 Orthomyxovirus (Influenza A) © Dr Linda Stannard,
University of Cape Town, South Africa
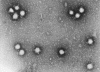 Figure 18 Bunyavirus From ICTV database
Figure 18 Bunyavirus From ICTV database
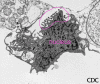 Figure 19b Vero E6
tissue culture cell infected with an arenavirus. Image shows
extracellular virus particles budding from the cell surface. Magnification
approx. 12,000 times. Image courtesy Cynthia Goldsmith, MS,
Infectious Disease Pathology Activity, DVRD, NCID, CDC
Figure 19b Vero E6
tissue culture cell infected with an arenavirus. Image shows
extracellular virus particles budding from the cell surface. Magnification
approx. 12,000 times. Image courtesy Cynthia Goldsmith, MS,
Infectious Disease Pathology Activity, DVRD, NCID, CDC |
SEGMENTED NEGATIVE STRAND
VIRUSES
Examples:
-
Orthomyxoviruses (figure 16 and
17)
-
Bunyaviruses
(include Hantavirus genus) (figure 18)
-
Arenaviruses (figure 19b)
ORTHOMYXOVIRUSES
(ORTHOMYXOVIRIDAE)
There are three groups of
influenza virus: A, B
and C. Influenza A virus is most intensively studied and influenza A
and B are the most important in human disease.
Influenza viruses are
pleomorphic virions (that is, they vary in shape). They have negative-sense, single-stranded RNA and
an
RNA genome that is SEGMENTED. There are eight RNA segments in influenza A. The
nucleocapsid is helical (figure 19). Virions contain RNA polymerase packaged within the virus particle
These viruses are enveloped and have two membrane glycoproteins (figure
19):
|
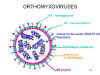 Figure 19 Structure of a typical orthomyxovirus
Figure 19 Structure of a typical orthomyxovirus |
Adsorption
and penetration
The virus adsorbs to receptors on the cell surface and is internalized by
endocytosis.
At acid pH of an endosome, HA undergoes a
conformational change and fusion occurs. Nucleocapsids are released to
cytoplasm.
|
 Figure 20
Figure 20
Transcription of orthomyxoviridae RNA |
Transcription, translation
and replication
Nucleocapsids are transported into the
nucleus. mRNA synthesis and replication of viral RNA occurs in the nucleus. This
is very unusual for an RNA virus. Influenza virus has an unusual
mechanism for acquiring a methylated, capped 5'end to its mRNAs.
A viral endonuclease (which is packaged in
the influenza virus) snips off the 5'end of a host capped, methylated mRNA
about 13-15 bases from the 5' end and uses this as a primer for viral mRNA
synthesis (figure 20) - hence all flu mRNAs have a short stretch at the 5' end which is
derived from host mRNA.
The viral RNA polymerase (transcriptase)
extends the primer and copies the template into complementary plus sense mRNA and adds a poly(A) tail.
Transcription results in 8 primary transcripts, one transcript per segment. Some segments give rise to
primary transcripts which can be alternatively spliced (since influenza virus
RNA synthesis occurs in the nucleus, it has access to splicing machinery), each
giving rise to two alternative transcripts. For example, the M segment gives
rise to two alternative mRNAs. These code for the M1 protein and the M2 protein.
Thus a
single segment can code for more than one protein since the virus has access to
splicing machinery. The mRNAs are translated in the
cytoplasm. Transmembrane proteins are moved to the plasma membrane while proteins needed for RNA
replication are transported to the nucleus.
|
| |
Replication of RNA
RNA replication occurs in the nucleus using a virus-coded enzyme (this may be same as the RNA polymerase involved in
transcription of mRNAs, or a modified version).
A full length, exact
complementary copy of virion RNA is made - this plus sense RNA is probably coated with
nucleocapsid protein as it is made.
Full length plus strand RNA is then
used as a template for full-length minus strand synthesis; again the new minus
strand is probably coated with nucleocapsid protein as it is made.
New minus strands can be used as
templates for replication, mRNA synthesis, or packaged.
Assembly
This occurs at the plasma membrane.
Nucleocapsids are transported out
of the nucleus while envelope proteins are transported
via the Golgi body to the plasma membrane.
The M1 protein interacts with both nucleocapsid and a modified region of the plasma membrane which
contains the glycoproteins HA and NA.
Virus then buds out through the host cell membrane.
Note:
-
HA needs to be cleaved before it can promote fusion.
Cleavage occurs as the virus leaves the cell or in the extracellular fluid. The
requirement for cleavage affects which tissues can produce infectious virus.
The cleaved protein needs to then undergo a conformational change, usually
caused by exposure to a acidic endosome environment when it infects the next
cell, before it can cause fusion.
-
NA probably helps the virus leave the cell
by removing sialic acid from receptors. NA may also help the virus penetrate mucus
to reach epithelial cells of the respiratory tract by enabling it to dissociate from
sialic acid-containing receptors in the mucus by destroying them. The
neuraminidase does not prevent the virus infecting new cells because
endocytosis is presumably faster than receptor removal.
There are similarities and
differences between the Paramyxovirus family and the Orthomyxovirus family,
members of both are enveloped, both contain negative sense, single stranded RNA,
have helical nucleocapsids. However, the two families are very different. There
is NO immunological relationship between the two families.
|
| |
|
PROPERTY
|
PARAMYXOVIRIDAE
|
ORTHOMYXOVIRIDAE
|
|
Genome
|
non-segmented
|
segmented
|
|
RNA synthesis
|
cytoplasmic
|
nuclear
|
|
Need for mRNA primer
|
no
|
yes
|
|
Hemagglutinin,neuraminidase
|
if both, part of same protein (HN)
|
Influenza A and B have both
but on 2 different proteins (HA and NA)
|
|
Syncytia formation
|
yes (F functions at at
normal physiol. pH)
|
no (HA functions at acid pH)
|
|
 Figure 21 Mammalian Reovirus Virion
Figure 21 Mammalian Reovirus Virion
The cryoEM data was from Tim Baker's Laborratory, Purdue
University. Movies were created by Stephan Spencer.
Copyright
1999 Dr Tim Baker and Stepthen M Spencer. From Dr J-Y Sgro's
Virusworld |
DOUBLE STRANDED RNA VIRUSES
REOVIRUS FAMILY
(REOVIRIDAE)
The Reovirus family include:
-
the members of the Reovirus genus
-
the members of the
Rotavirus genus
-
the members of the Orbivirus genus
(e.g. Bluetongue virus)
-
the members of the
Coltivirus family (e.g. Colorado tick fever virus)
|
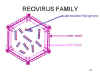 Figure 22 Structure of a typical reovirus Adapted from Joklik et al. Zinsser
Microbiology 20th Ed.
Figure 22 Structure of a typical reovirus Adapted from Joklik et al. Zinsser
Microbiology 20th Ed. |
Reoviruses have icosahedral symmetry
and a multiple
layered capsid (inner and outer capsid) (figure 22)
The RNA is double stranded. There are 10-12 segments (depending on the genus of the Reovirus family
to which the virus belongs) (figure 22).
There are some significant
differences in the life cycle of members of the reovirus family and of the
rotavirus family. Due to their clinical importance in humans, we shall focus on
rotaviruses.
|
 Figure 23 Rotavirus (A double-capsid particle (left), and a single, inner, capsid (right))
Copyright Dr
Linda
Stannard, University of Cape Town, South Africa
Figure 23 Rotavirus (A double-capsid particle (left), and a single, inner, capsid (right))
Copyright Dr
Linda
Stannard, University of Cape Town, South Africa |
ROTAVIRUSES
(rota = wheel (from appearance of virions in the electron-microscope)) (figure 23)
Adsorption, penetration
and uncoating
It is still not clear what exactly what
happens in vivo during the entry of
rotaviruses into the cell.
There appears to be a need for a protease to remove some of the outer
layer of the capsid and to
generate an "intermediate sub-viral particle" (ISVP) before the virus can enter
the cytoplasm. In vivo, the ISVPs are probably
generated by protease digestion in the GI tract.
A viral attachment protein is then exposed on the ISVP,
probably at the vertices, and binds to host cell receptors. The activated ISVP enters the
cytoplasm directly or via endocytosis. In the cytoplasm, the virion RNA is
copied by the viral RNA polymerase while still in a nucleocapsid that has fewer
proteins associated with it than are associated with the ISVP or the virion.
Transcription and
translation
Double stranded RNA does not
function as an mRNA and so the initial step is to make mRNA (transcription).
The mRNAs are made by virally-coded RNA polymerase packaged in the virion. The
RNA is capped and methylated by virion packaged
enzymes. It is then
extruded from the vertices of the capsid.
|
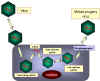 Figure 24
Figure 24
Replication of reoviridae |
The mRNAs are translated and the resulting
viral proteins assemble to form an immature capsid. The mRNAs are packaged into
the immature capsid and are then copied within the capsid to form double
stranded RNAs (It is not known how the
virus ensures that each particle gets
one copy of the 11 different mRNAs) (figure 24). More mRNAs are now made by the
newly formed immature capsids.
Assembly
More proteins are made and eventually the
immature capsids bud into the lumen of the endoplasmic reticulum. In doing so,
they acquire a transient envelope which is lost as they mature. This is a very
odd feature of the rotaviruses.
Release
This probably occurs via cell lysis.
Note: The entire replication cycle occurs in the cytoplasm
|
|
 Return to the Virology section of Microbiology and Immunology On-line
Return to the Virology section of Microbiology and Immunology On-line
 Return to the Home Page of Microbiology and Immunology On-line
Return to the Home Page of Microbiology and Immunology On-line
This page last changed on
Tuesday, May 31, 2016
Page maintained by
Richard Hunt
|

 Figure 1 Polio virus © J-Y Sgro, Used with permission.
From
Virus World
Figure 1 Polio virus © J-Y Sgro, Used with permission.
From
Virus World
 Figure 4 Adapted from Schaechter et al., Mechanisms of
Microbial Disease, 2nd Ed.
Figure 4 Adapted from Schaechter et al., Mechanisms of
Microbial Disease, 2nd Ed. Figure 6 Rhabdovirus on a Fish Epithelial Cell
©
Dennis Kunkel Microscopy, Inc.
Used with permission
Figure 6 Rhabdovirus on a Fish Epithelial Cell
©
Dennis Kunkel Microscopy, Inc.
Used with permission  Figure 7 Structure of a typical rhabdovirus
Figure 7 Structure of a typical rhabdovirus Figure 8
Figure 8 Figure 9
Figure 9
 Figure 10
Figure 10
 Figure 11 Paramyxovirus ©
Dr
Linda
Stannard,
University of Cape Town, South Africa
(used with permission)
Figure 11 Paramyxovirus ©
Dr
Linda
Stannard,
University of Cape Town, South Africa
(used with permission)
 Figure 12 Structure of a typical paramyxovirus
Figure 12 Structure of a typical paramyxovirus Figure 13
Figure 13 Figure 14
Figure 14 Figure 15
Figure 15 Figure 19 Structure of a typical orthomyxovirus
Figure 19 Structure of a typical orthomyxovirus Figure 20
Figure 20 Figure 21 Mammalian Reovirus Virion
Figure 21 Mammalian Reovirus Virion  Figure 22 Structure of a typical reovirus Adapted from Joklik et al. Zinsser
Microbiology 20th Ed.
Figure 22 Structure of a typical reovirus Adapted from Joklik et al. Zinsser
Microbiology 20th Ed. Figure 23 Rotavirus (A double-capsid particle (left), and a single, inner, capsid (right))
Copyright Dr
Linda
Stannard, University of Cape Town, South Africa
Figure 23 Rotavirus (A double-capsid particle (left), and a single, inner, capsid (right))
Copyright Dr
Linda
Stannard, University of Cape Town, South Africa Figure 24
Figure 24
 Structure of Polio Type 1 Mahoney. X-ray data from Hogle et al.(Harvard Univ.), PDB entry 2PLV, rendered with GRASP (A.Nicholls, Columbia Univ.). Courtesy of
Structure of Polio Type 1 Mahoney. X-ray data from Hogle et al.(Harvard Univ.), PDB entry 2PLV, rendered with GRASP (A.Nicholls, Columbia Univ.). Courtesy of





